The UK has formed a genome sequencing alliance, to enable rapid, broad, large-scale sequencing analysis of samples from patients testing positive for COVID-19. The network aims to sequence the virus from every patient sample that has tested positive. The resulting data will help deliver insights into how the virus is transmitted and how it evolves.
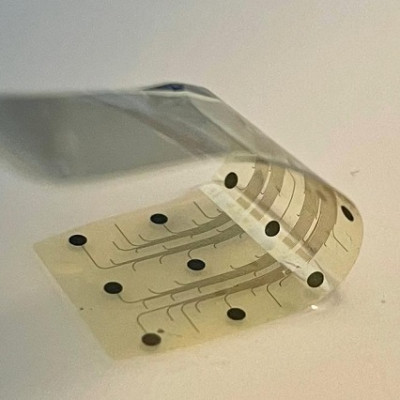
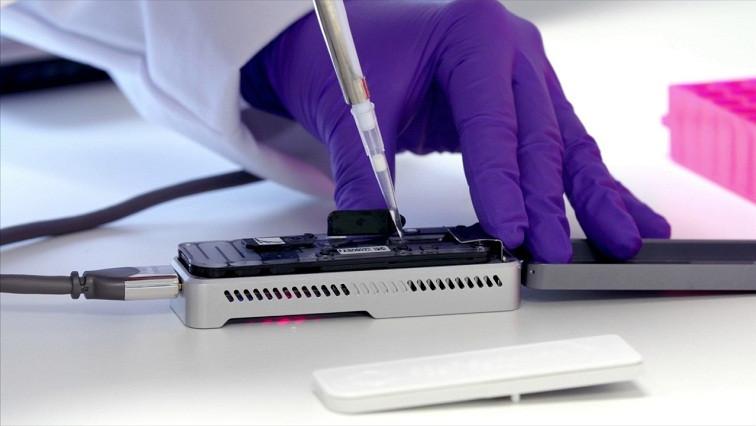
Oxford Nanopore is delighted to support participating teams across the UK in this project, including in the cities of Birmingham, London, Edinburgh, Glasgow, Nottingham, Sheffield, Liverpool, Cardiff, Exeter and Cambridge. Nanopore sequencing can deliver a whole genome sequence of the virus in as little as 7 hours, on the portable MinION or desktop GridION devices.
The ARTIC network has been key to enabling rapid sequencing of the SARS-CoV-2 virus in the UK and globally. Since releasing methods that were compatible with nanopore sequencing in January 2020, the ARTIC protocol has been used for rapid sequence generation, originally in China and now all over the world.
Oxford Nanopore’s MinION device has also been used in other viral outbreaks including Lassa Fever, Swine flu, yellow fever, and in outbreaks caused by bacterial or fungal pathogens. ARTIC’s previous development of rapid protocols and methods for viral sequencing have previously been used in multiple outbreaks including Zika in Brazil and Ebola in Guinea/DRC.
Oxford Nanopore is supporting many groups around the world in their COVID-19 genomics. A range of work is underway including whole genome sequencing, metagenomic analysis and direct RNA sequencing. Read more about the UK announcement here.
Read the original article on Oxford Nanopore Technologies.