Ilaria Testa, an associate professor at KTH and researcher at the Science for Life Laboratory, says the technique is capable of producing images with precision that until now has been exclusive to electron microscopy. The advance was reported in Nature Biotechnology, with near-molecular scale images of proteins in the brain’s hippocampal neurons.
“This technique allows us to image proteins with a completely new level of 3D spatial details, and importantly in situ within the cell,” she says.
Dubbed 3D pRESOLFT, the KTH technique can visualize proteins on a wider scale than possible with electron microscopy; and it can do so without killing the cell and sectioning it – two necessary steps with an electron microscope.
“Here, we don’t need to do that. The cells are happily moving and undergoing important functions,” Ilaria says.
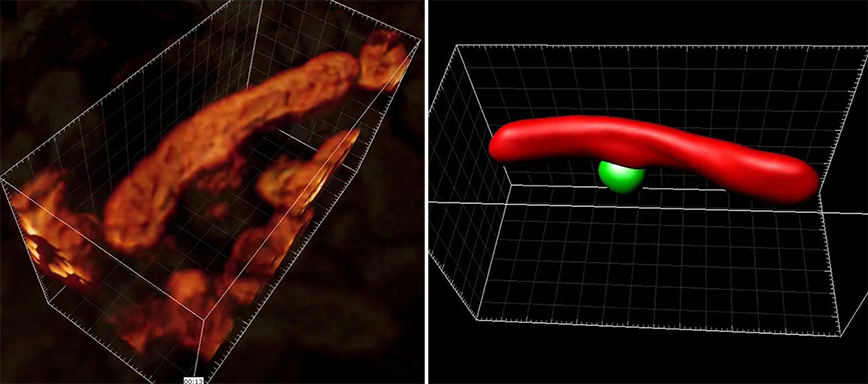

The researchers generated this 3D image of living mitochondria, left, in a human cell. Their new method enables disentangling of a mitochondrial network and optically “slicing” through an organelle, capturing multiple images that can be reconstructed in 3D. In this case, the rendering on the right reveals a small, round mitochondrion with a longer, larger one connect to each other.
Ilaria’s lab developed the technique as part of its focus on investigating the localization and function of neuronal proteins, especially in synapses and axons, where trafficking organelles and protein complexes are so crowded that they need to be visualized in high resolution in order to be examined.
In conventional fluorescence microscopy, visible light is used to illuminate cells and tissues that are colored with a fluorescent dye – but the method is limited to creating 2D images, typically at low resolution. 3D pRESOLFT improves upon the technique by using a combination of interference patterns involving switchable fluorescence dye that can be turned on and off, like a light switch, while a large volume of parallel images are recorded. The sample is exposed to less light overall, preventing the sample from fading.
“Because the light is gentle we can look at living cells but with a new precision, that is, down to 50 nm in scale, or 20,000 times smaller than a human hair,” she says.
The ability to view living cells in 3D with such precision makes it possible to study how proteins take part in important and yet poorly understood physiological processes, she says.
“We now can see the 3D architecture of brain cells, and examine molecules that we consider important for learning and memory formation – and understand how they change location and shape when exposed to certain stimulations,” she says.
Read the original article on KTH Royal Institute of Technology.