Several processes in the human body are regulated by biochemical reactions involving hydrogen peroxide (H2O2). Although it can act as a ‘secondary messenger’, relaying or amplifying certain signals between cells, H2O2 is generally toxic because of its oxidant character. The latter means that it converts (oxidizes) biochemical molecules like proteins and DNA. The oxidizing property of H2O2 is of potential therapeutic relevance for cancer, though: deliberately causing tumor cells to increase their H2O2 concentration would be a way to destroy them. In light of this, but also for monitoring pathologies associated with H2O2 overproduction, it is crucial to have a means to reliably quantify hydrogen peroxide concentrations in the extracellular environment. Now, Leonardo Puppulin from Nano Life Science Institute (WPI-NanoLSI), Kanazawa University and colleagues have developed a sensor for measuring concentrations of H2O2 in the vicinity of cell membranes, with nanometer-resolution.
The biosensor consists of a gold nanoparticle with organic molecules attached to it. The whole cluster is designed so that it anchors easily to the outside of a cell’s membrane, which is exactly where the hydrogen peroxide molecules to be detected are. As attachment molecules, the scientists used a compound called 4MPBE, known to have a strong Raman scattering response: when irradiated by a laser, the molecules consume some of the laser light’s energy. By measuring the frequency change of the laser light, and plotting the signal strength as a function of this change, a unique spectrum is obtained — a signature of the 4MPBE molecules. When a 4MPBE molecule reacts with a H2O2 molecule, its Raman spectrum changes. Based on this principle, by comparing Raman spectra, Puppulin and colleagues were able to obtain an estimate of the H2O2 concentration near the biosensor.
After developing a calibration procedure for their nanosensor — relating the H2O2 concentration to a change in Raman spectrum in a quantitative way is not straightforward — the scientists were able to produce a concentration map with a resolution of about 700 nm for lung cancer cell samples. Finally, they also succeeded in extending their technique to obtain measurements of the H2O2 concentration variation across cell membranes.
Puppulin and colleagues conclude that their “novel approach may be useful for the study of actual H2O2 concentrations involved in cell proliferation or death, which are fundamental to fully elucidate physiological processes and to design new therapeutic strategies.”
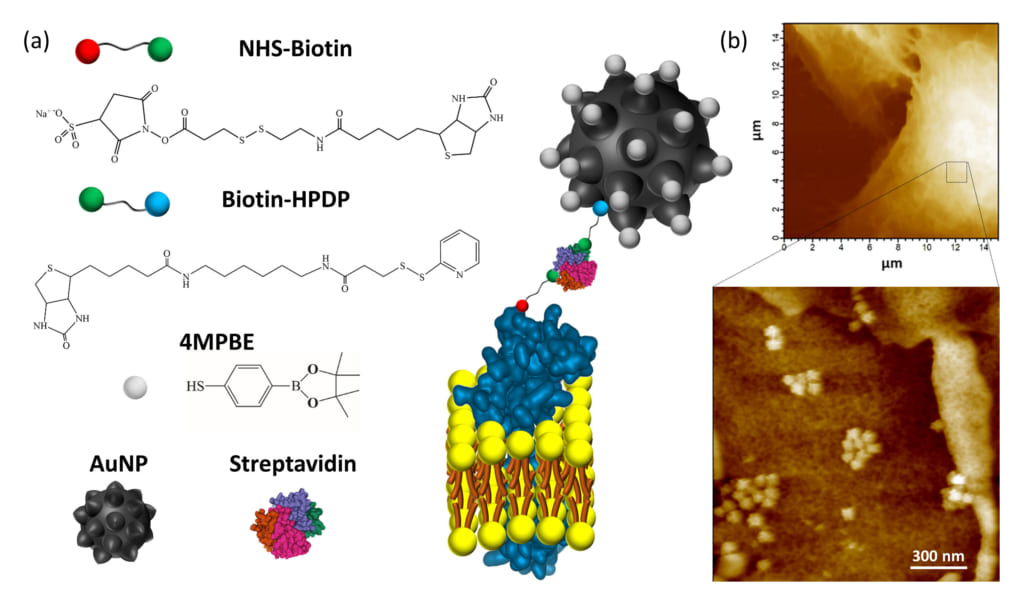
Figure1. (a) Explanatory sketch of the plasma membrane anchored nanosensor. The compounds used for gold nanoparticles (AuNP) conjugation are the H2O2-sensitive 4-mercaptophenylboronic pinacol ester (4MPBE) and Biotin-HPDP. Biotinylation of the plasma membrane protein ectodomains is obtained using NHS-Biotin. The anchoring of the conjugated AuNP and NHS-Biotin is given by Streptavidin reacting with the two biotin moieties.
(b) AFM analysis was performed on A549 lung cancer cells after nanosensor anchoring and fixation. High resolution AFM images confirmed the presence of the nanosensor, which is in contact with the cell surface and is capable of detecting endogenous H2O2 in a very shallow region (i.e., 90 nm) of the extracellular fluid in contact with the plasma membrane.
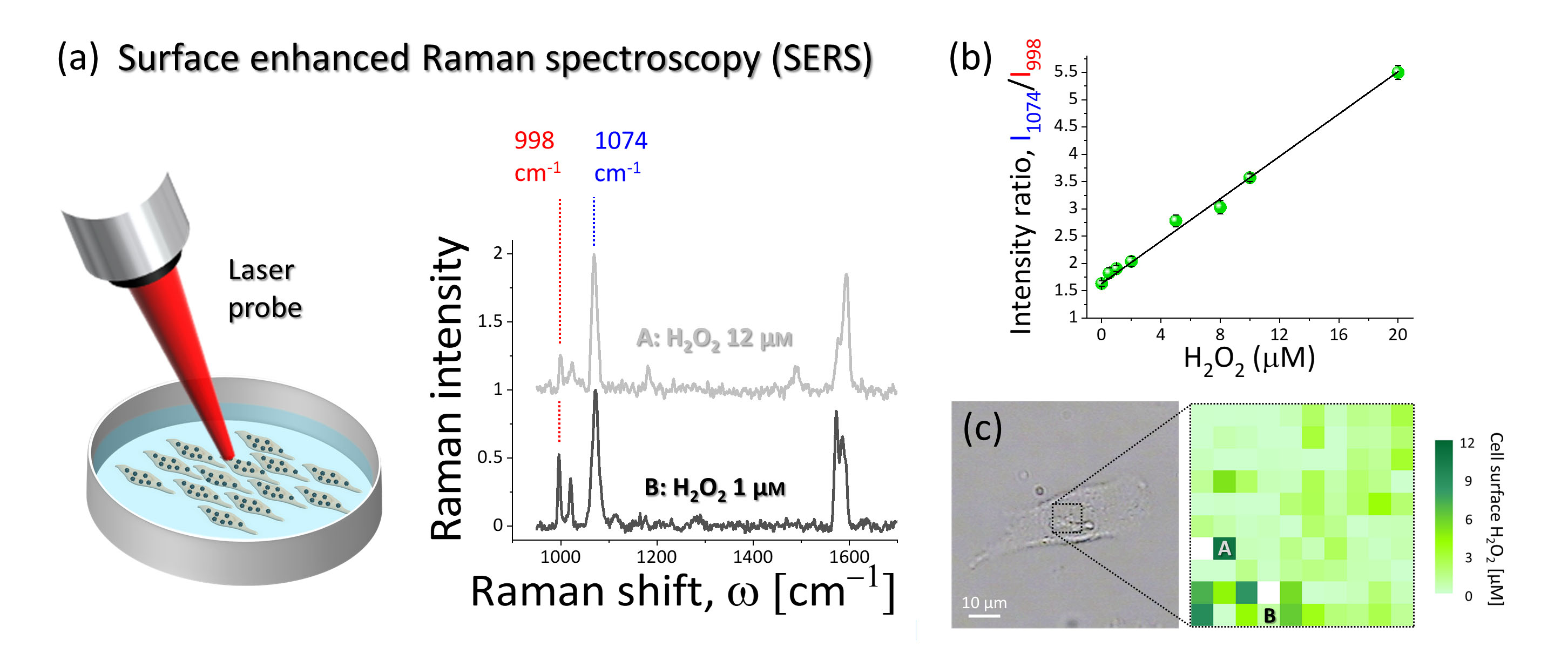
Figure2. (a) Surface enhanced Raman spectroscopy was used to detect and quantify the H2O2-induced modification of 4MPBE molecules assembled on the gold surface of the nanosensor anchored to the cells. The intensity of the Raman band at 998 cm-1 depends on H2O2 concentration, while the band at 1074 cm-1 did not show modification and it can be used for normalization of the spectral lines.
(b) Results of the nanosensor calibration. The ratio of the intensity at 1074 cm-1 to the intensity at 998 cm-1 showed linear dependence with respect to H2O2 concentration.
(c) Example of SERS hyperspectral map of extracellular endogenous H2O2 collected from the surface of the A549 cell shown in the bright-field image. The SERS spectra shown in (a) were collected from the locations A and B.
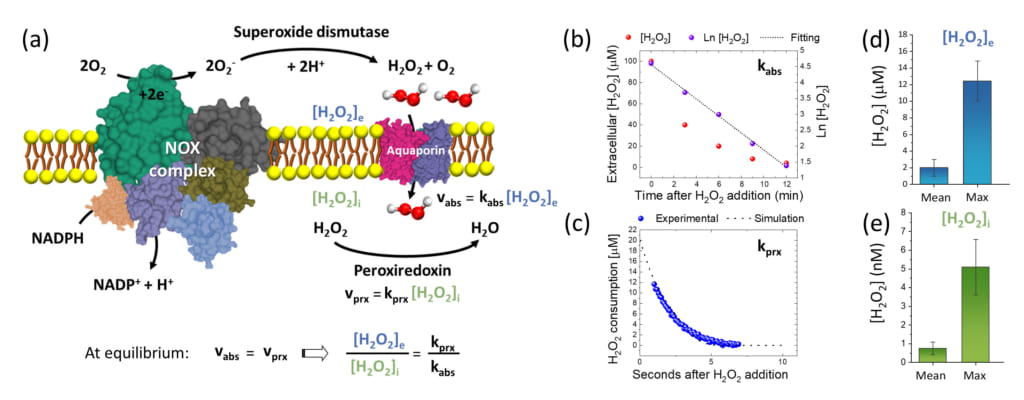
Figure 3. (a) Explanatory schematic of the extracellular production of H2O2 by NOX complex, the consequent absorption through Aquaporin channel and the reaction with intracellular peroxiredoxin. At steady-state condition, the concentration ratio of extracellular H2O2 to intracellular H2O2 can be estimated by the ratio of the rate constant of H2O2 reduction by peroxiredoxin (kprx) to the rate constant of H2O2 absorption through the plasma membrane (kabs).
(b)-(c) Typical results of redox biology experiments with fitting lines from which we measured kabs and kprx, respectively.
(d) Mean surface extracellular [H2O2]e and the typical maximum of [H2O2]e measured on A549 cells using the newly designed nanosensor. (e) Mean intracellular [H2O2]i and the typical maximum of [H2O2]i estimated on A549 cells according to the model reported in (a) and using the results from (b)-(d).
[Background]
Surface-enhance Raman spectroscopy
The biosensor developed by Leonardo Puppulin from Kanazawa University and colleagues is based on a method called surface-enhanced Raman spectroscopy (SERS). The principle derives from Raman spectroscopy, in which differences between the incoming and the outcoming frequencies of laser light irradiated onto a sample are analyzed. The spectrum obtained by plotting the signal strength as a function of frequency difference is characteristic for the sample, which can in principle be a single molecule. Typically, however, the signal coming from one molecule is too weak to detect, but the effect can be enhanced when the molecule is absorbed on a rough metal surface. Puppulin and colleagues applied the technique to (indirectly) detect hydrogen peroxide; their Raman-responsive molecule is a compound called 4MPBE, which is modified when exposed to hydrogen peroxide.
Read the original article on Kanazawa University.