Viruses kill millions around the world each year. "In addition to the novel coronavirus, leading viral killers include hepatitis, HIV, HPV," said Lela Vukovic, Assistant Professor of Chemistry, University of Texas at El Paso.
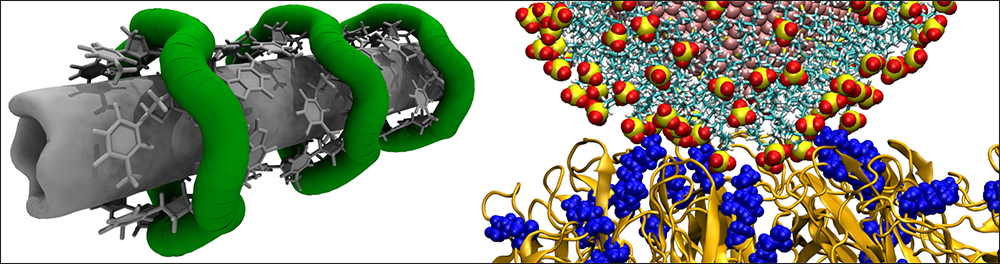
Vukovic's research combines molecular dynamics simulations with machine learning to better understand and engineer nanoscale systems for medicine.
Researchers are constantly trying to figure out new therapeutics that will help prevent infection or act therapeutically to reduce symptoms for one virus at a time. "Another strategy," Vukovic said, "would be to find therapies that are broad spectrum and simultaneously act on a number of different viruses."
Many viral infections start with the virus binding to heparan sulfate molecules on the host cell's surface. Working with experimentalists led by Francesco Stellacci of the Swiss Federal Institute of Technology Lausanne (EPFL), and in collaboration with Petr Král at the University of Illinois at Chicago, Vukovic helped to investigate nanoparticles with solid cores and ligands attached that mimic the heparan sulfate molecules and their microscopic action on several viruses.
They found that nanoparticles with certain ligands can attach to the viruses, which soon after may disintegrate.
"Such virus-destructing materials can be prepared," Vukovic said at a recent seminar at the Texas Advanced Computing Center (TACC). "The question is: Are there hints we can get from computational modeling to design new, better materials and understand the mechanism that causes the virus capsid to break?"
Since nanoparticles are minute, they can't be imaged clearly at the atomic level and microsecond timescales at which the reactions happen. So Vukovic created models of the atomic structure of virus, as well as the nanoparticles with ligands of various lengths attached.
Using TACC supercomputers, she simulated how the viral proteins and nanoparticles interact with each other. She found that the virus binds and makes numerous contacts with longer ligands.
Not only that. The nanoparticles bind at the junction of two proteins and, like a wedge, increase the distance between viral proteins, breaking the contacts and disintegrating the virus. The initial findings research were published in Nature Materials in 2018, and new results, obtained by the student Parth Chaturvedi, have been posted on bioRxiv (August 2021).
NUANCED DESIGNS OF NANOSENSORS
Vukovic's interest in modeling nanoparticles for medicine led her to her next project, helping to design nanosensors that are small, fast, and sensitive enough to detect microscopic amounts of neurotransmitters in the brain.
The basis of the technology are carbon nanotubes — cylinders 10,000 times narrower than the average human hair — which have found applications in various fields, including electronics, optics, and most recently medicine.
Carbon nanotubes, or CNTs, researchers found, have an unusual property. They can spontaneously luminesce in certain circumstances with a light that can be detected outside the body. However, they can't operate in the body without modification.
One approach that has proven successful involves wrapping the CNT in DNA. The Landry lab at the University of California, Berkeley were experimenting with DNA strands of various lengths and makeups to see whether the CNT gave off a strong light emission when exposed to dopamine, and were getting mixed results.
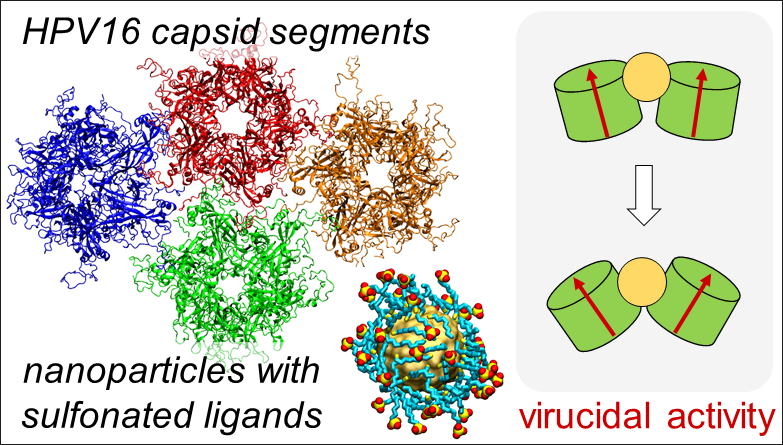
Solid core nanoparticles coated with sulfonated ligands that mimic heparan sulfate proteoglycans can exhibit virucidal activity against many viruses. Vukovic's computational research explores the mechanism for this activity to enable better broad-spectrum antivirals.
"The screening approach works, but it doesn't provide a good understanding of why it works or how to design it better in the future. Can we do something more systematic?" Vukovic asked.
She undertook a series of computational experiments on Stampede2, TACC's leading supercomputer at the time, exploring the 3D structure, energy landscape, and binding patterns of CNTs wrapped with DNA.
She and her student Ali Alizadehmojarad found that DNA of certain lengths wrap around the nanotube like a ring, while others wrap it as a helix or irregularly. These different binding patterns lead to different luminescence in the presence of neurotransmitters. The ring-wrapped CNT of one type of DNA, she and the Landry lab found, was far more effective at detecting and signaling the presence of neurotransmitters. The research was published in a series of papers in Nano Letters in 2018 and Advanced Material Interfaces in 2020.
NANO-PIVOT
The challenges, and achievements of the sensor project, inspired an epiphany in Vukovic.
She had successfully explored the experimental mysteries of CNTs at the atomic level using molecular dynamics simulations and provided critical insights. "But I'm only doing one molecule at a time," Vukovic said. "As a theoretician, what can I contribute? If I test 10 molecules, I don't even scratch the surface."
Her realization led her to incorporate AI and data-driven methods into her approach. "We completely switched our research; learned new methods. For the last two years, we've been working on that."
This period of growth and learning led Vukovic and her team, Payam Kelich and Huanhuan Zhao, to their most recent project: working with the Landry lab on the discovery of new optical sensors made of DNA-CNT conjugates to detect the serotonin molecule. As a key molecule that impacts our mood and happiness, there is a great interest in detecting serotonin presence and amounts in different body tissues.
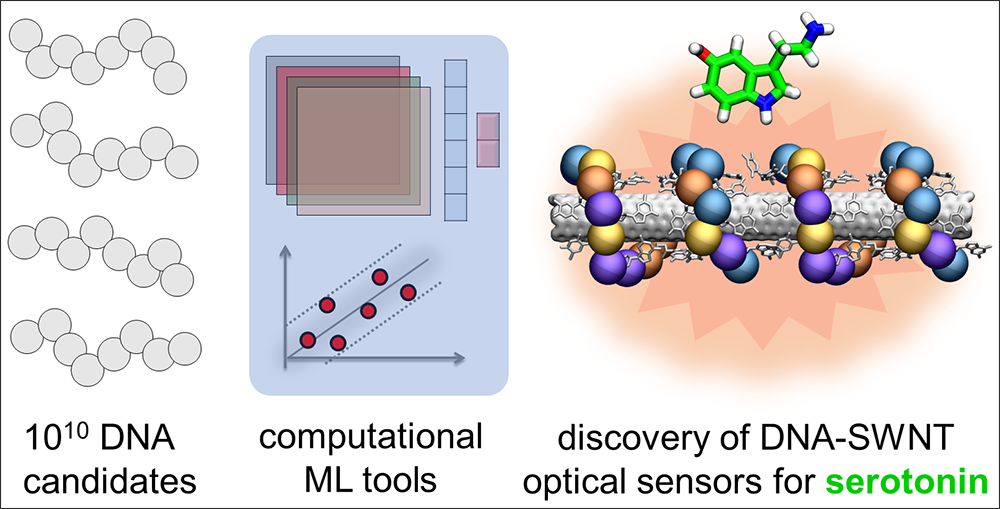
Near-infrared (nIR) fluorescence response datasets for ~100 DNA-single walled nanotube conjugates were used to train machine learning (ML) models to predict new unique DNA sequences with strong optical response to neurotransmitter serotonin.
Recently, Vukovic lab developed new AI-based computational tools that train models to learn from Landry's experimental data and predict new sensors of serotonin.
The collaboration is bearing fruit. A first paper, just posted on bioRxiv (August 2021), described efforts to computationally predict new serotonin sensors and experimentally validate the predictions. So far, the approach led to discovery of five new serotonin DNA-CNT sensors with a higher response than observed in previous sensors. (This research is supported by a new grant from the National Science Foundation.)
Vukovic is able to tackle these massive and ambitious computational challenges in part because of her access to some of the most cutting-edge scientific instruments on the planet through the University of Texas Research Cyberinfrastructure (UTRC) program. Started in 2010, the initiative provides powerful computing and data resources at no cost to Texan scientists, engineers, students, and scholars at all 13 UT System institutions.
"None of these projects would have been possible without TACC," Vukovic said. "When we were ready to run, we were given the time we needed and were able to advance quickly and get things done."
As a computational chemist, Vukovic says she is trying to use her knowledge to contribute to practical applications in medicine and beyond. "We are thinking deeply about how to contribute and working on projects where computing can make a real difference."
Read the original article on Texas Advanced Computing Center, The University of Texas at Austin.