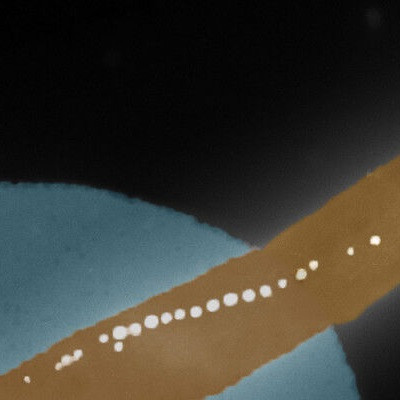
Even though proteins are crucial to life – providing you with vision and neural connections to read this text, for example – the way they change shape is still poorly understood. As published on Monday 30 August in Nature Nanotechnology, a team led by Cees Dekker at Delft University of Technology developed a new technique, called the Nanopore Electro-Osmotic trap (NEOtrap), to study individual protein molecules for much longer than was possible before. The NEOtrap allows the researchers to measure how single proteins change their shape over time.
Like a cork on a bottle
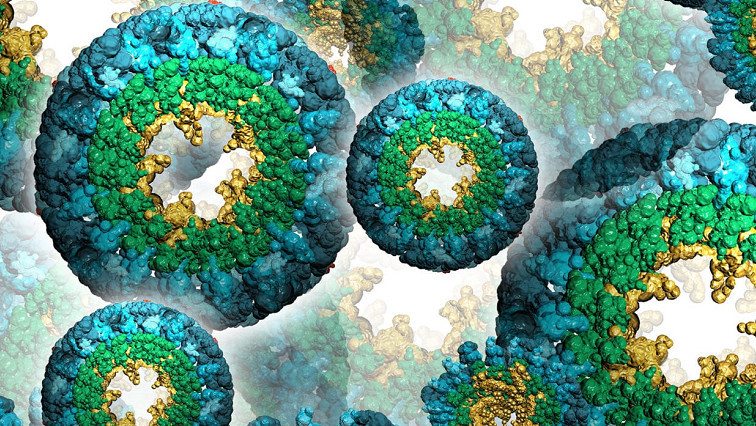
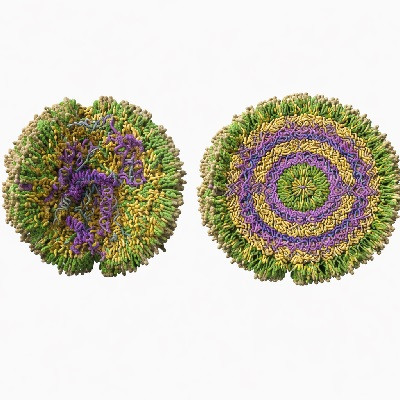
The NEOtrap combines two nanotechnologies: solid-state nanopores and DNA origami. Nanopores are small holes that scientists use as sensors for single molecules such as proteins. Since proteins normally pass through the small hole in microseconds, they can be recorded only briefly. By sealing the nanohole with a nanoscale ball built entirely from DNA (!), the researchers can lock the protein in place for hours, much like a cork seals a bottle of wine. Hendrik Dietz and his group at Technical University of Munich built this nano-ball using an approach called ‘DNA origami’ – a technique that mimics macroscale origami folding – using nanoscale DNA strands rather than paper.
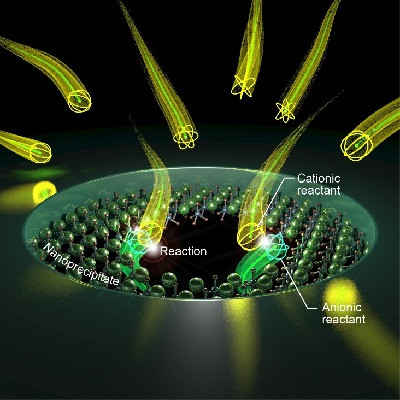
Lead author of the paper Sonja Schmid, who developed the NEOtrap as a postdoc in Dekker’s lab, explains: ‘This DNA-origami nano-ball acts like a sponge that sucks water through the nanopore, attracting a single protein to the nanopore and trapping it there. This means we can study that protein for very long periods of time. In this work we already demonstrate that we can distinguish between different types of proteins, and even different functional shapes of one and the same protein.’
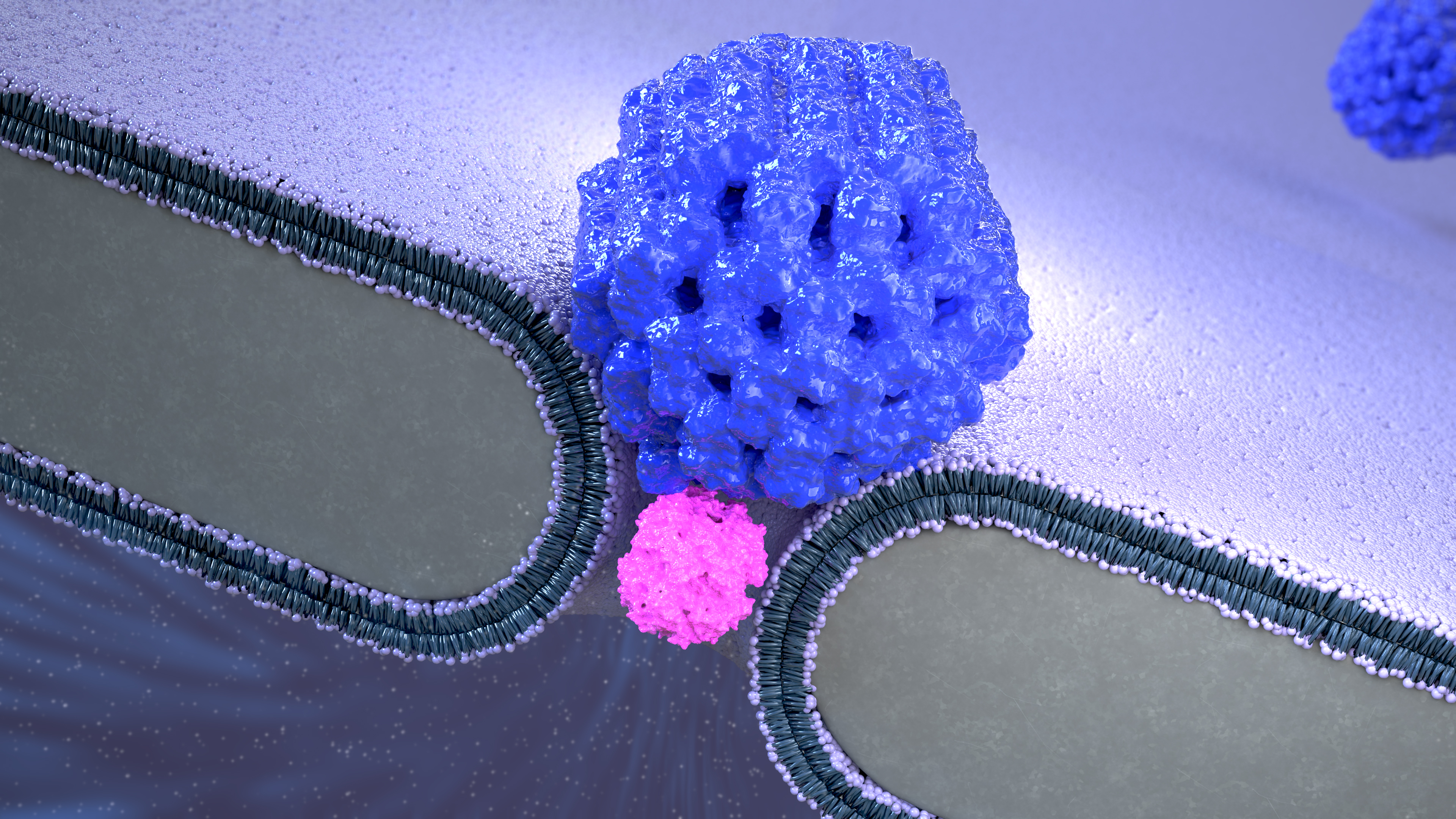
Artist impression of the Nanopore Electro-Osmotic trap (NEOtrap) – a new method for analyzing individual molecules. A DNA-origami nanoball covers a solid nanohole, allowing it to trap individual unmodified protein molecules, enabling hour-long conformation-sensitive analyses.
Radical advancement in the field
Cees Dekker adds: ‘This new technique really is a major step forward – an anonymous reviewer of our paper called it “one of the most radical advancements in the nanopore sensing field”. Notably, the NEOtrap enables us to trap a single native protein without the need to modify the molecule of interest, as opposed to previous techniques. This technique can, for example, help researchers to uncover the underlying mechanism of enzymes and other important proteins that change their shape to facilitate chemical reactions.’
The NEOtrap enables scientists around the globe to run entirely new experiments, with the potential to reveal previously overlooked functional features of proteins and hence spark innovation in biomedicine, biotechnology, and more. Schmid (who now started her own lab at Wageningen) and Dekker are planning many follow up studies of the dynamics of single proteins in the years to come.
Read the original article on TU Delft.