Self-assembly, introduced in the early 2000s, gives scientists a means to “pre-program” particles, allowing for the building of materials without further human intervention—the microscopic equivalent of Ikea furniture that can assemble itself.
The breakthrough, reported in the journal Nature, centers on emulsions—droplets of oil immersed in water—and their use in the self-assembly of foldamers, which are unique shapes that can be theoretically predicted from the sequence of droplet interactions.
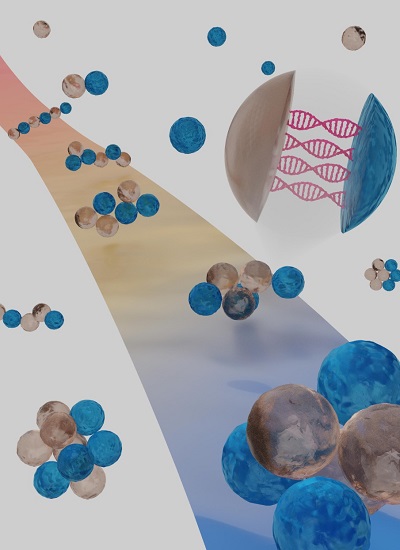

The illustration shows how droplets with different DNA strands first combine into chains, which are then programmed to fold into specific geometries, analogous to protein folding. The carpet highlights one folding pathway of a hexamer chain folding into a polytetrahedron. The zoom shows how the formation of DNA double helices drives droplet-droplet binding.
The self-assembly process borrows from the field of biology, mimicking the folding of proteins and RNA using colloids. In the Nature work, the researchers created tiny, oil-based droplets in water, possessing an array of DNA sequences that served as assembly “instructions.” These droplets first assemble into flexible chains and then sequentially collapse, or fold, via sticky DNA molecules. This folding yields a dozen types of foldamers, and further specificity could encode more than half of 600 possible geometric shapes.
“Being able to pre-program colloidal architectures gives us the means to create materials with intricate and innovative properties,” explains Jasna Brujic, a professor in New York University’s Department of Physics and one of the researchers. “Our work shows how hundreds of self-assembled geometries can be uniquely created, offering new possibilities for the creation of the next generation of materials.”
The research also included Angus McMullen, a postdoctoral fellow in NYU’s Department of Physics, as well as Maitane Muñoz Basagoiti and Zorana Zeravcic of ESPCI Paris.
The scientists emphasize the counterintuitive, and pioneering, aspect of the method: Rather than requiring a large number of building blocks to encode precise shapes, its folding technique means only a few are necessary because each block can adopt a variety of forms—as seen in the below images.

Microscopy images show a chain of alternating blue and yellow droplets folding into a crown geometry through blue-blue, blue-yellow, and finally yellow-yellow interactions, mediated by sticky DNA strands. Microscopic droplets are programmed to interact via sticky DNA strands to uniquely fold into well-defined shapes, as shown here.
“Unlike a jigsaw puzzle, in which every piece is different, our process uses only two types of particles, which greatly reduces the variety of building blocks needed to encode a particular shape,” explains Brujic. “The innovation lies in using folding similar to the way that proteins do, but on a length scale 1,000 times bigger—about one-tenth the width of a strand of hair. These particles first bind together to make a chain, which then folds according to preprogrammed interactions that guide the chain through complex pathways into a unique geometry.”
“The ability to obtain a lexicon of shapes opens the path to further assembly into larger scale materials, just as proteins hierarchically aggregate to build cellular compartments in biology,” she adds.
Images depicting this process and resulting shapes may be downloaded from Google Drive.
The work was supported by grants from the National Science Foundation (DMR-1420073, PHY17-48958, DMR-1710163) as well as by the Paris Region under the Blaise Pascal International Chairs of Excellence.
Read the original article on New York University (NYU).