The method, based on nanopore-based technology, was reported in an article in Frontiers in Cellular and Infection Microbiology.
Immunocompromised cancer patients are susceptible to various microbial infections and have a higher mortality rate than patients with other infections. Therefore, early identification of microorganisms is essential for the implementation of anti-infective therapy in patients.
In this study, the researchers collected samples from 56 immunocompromised cancer patients with suspicious infections. The samples included blood and sputum.
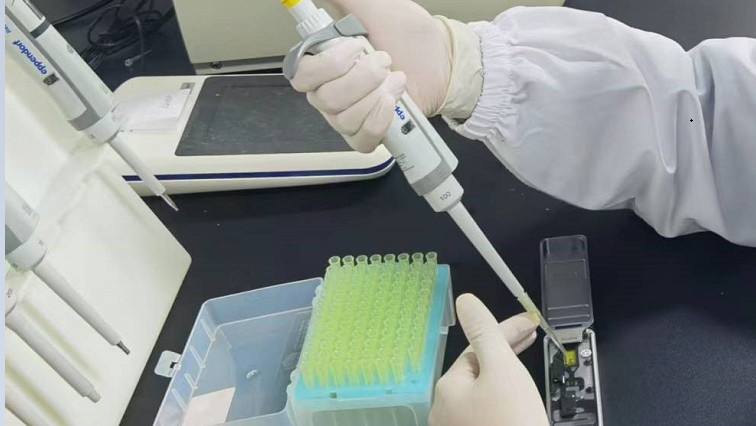
They isolated microbes and generated amplicon-based sequencing libraries, and sequenced the libraries utilizing the Oxford Nanopore MinION, a 90 g portable sequencer. Nanopore sequencing showed a significantly higher sensitivity than that of the conventional culture method. This advantage existed both in blood samples and non-blood samples.
Compared with traditional method, nanopore amplicon sequencing method showed more samples with bacterial infections, infections from fastidious pathogens, and co-infections. The mean turnaround time was approximately 17.5 hours, much shorter than the conventional culture assay.
"This method is rapid and accurate," said GU Hongcang, first author of the study, "it can detect pathogens among immunocompromised cancer patients with suspected infections."
This novel and high-sensitive method would improve the prognosis of cancer patients with low immune function by facilitating the prompt diagnosis of infections and the implementation of timely and accurate anti-infective treatments.
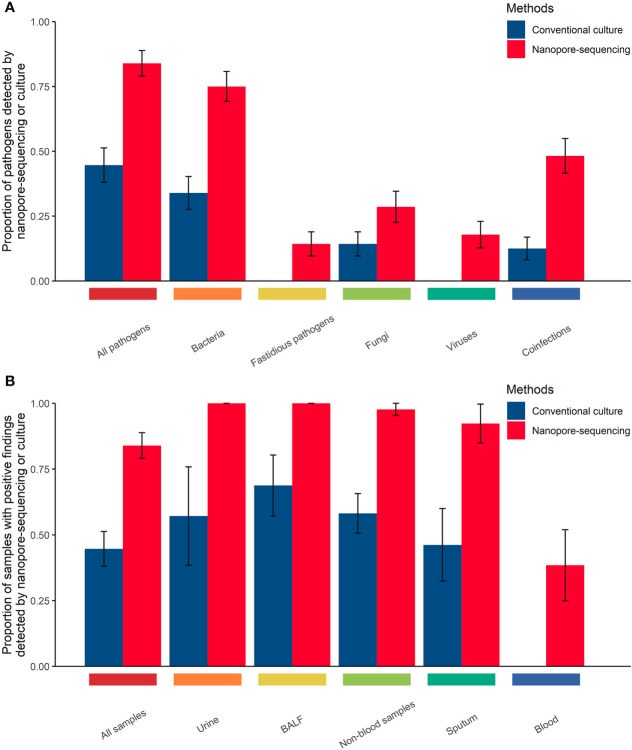
Comparison of the performance of those two methods in detecting pathogens in body fluid samples from immunocompromised cancer patients with suspected infections.
Read the original article on Chinese Academy of Sciences (CAS).