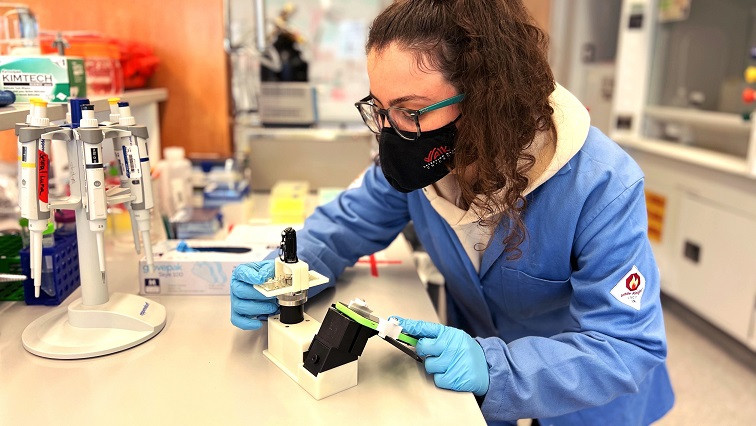
“One of our big innovations, beyond getting the DNA motors to perform logic computations, is finding a way to convert that information into a simple output signal — motion or no motion,” says Selma Piranej, an Emory University PhD candidate in chemistry, and first author of the paper. “This signal can be read by anyone holding a cell phone equipped with an inexpensive magnifying attachment.”
“Selma’s breakthrough removes major roadblocks that stood in the way of making DNA computers useful and practical for a range of biomedical applications,” says Khalid Salaita, senior author of the paper and an Emory professor of chemistry at Emory University. Salaita is also on the faculty of the Wallace H. Coulter Department of Biomedical Engineering, a joint program of GeorgiaTech and Emory.
The motors can sense chemical information in their environment, process that information, and then respond accordingly, mimicking some basic properties of living cells.
“Previous DNA computers did not have directed motion built in,” Salaita says. “But to get more sophisticated operations, you need to combine both computation and directed motion. Our DNA computers are essentially autonomous robots with sensing capabilities that determine whether they move or not.”
The motors can be programmed to respond to a specific pathogen or DNA sequence, making them a potential technology for medical testing and diagnostics. Another key advance is that each motor can operate independently, under different programs, while deployed as a group. That opens the door for a single massive array of the micron-sized motors to carry out a variety of tasks and perform motor-to-motor communication.
“The ability for the DNA motors to communicate with one another is a step towards producing the kind of complex, collective action generated by swarms of ants or bacteria,” Salaita says. “It could even lead to emergent properties.”
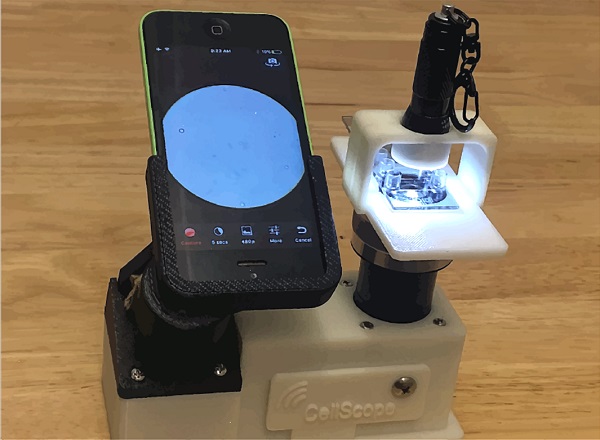

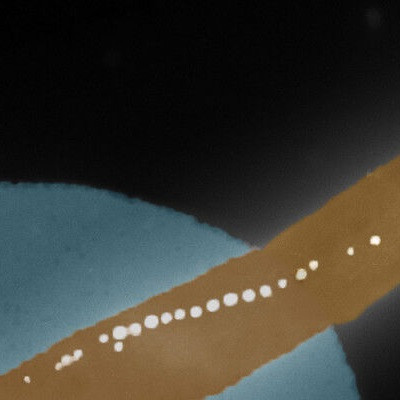
The high speed of the rolling DNA-based motor allows a simple smart phone microscope to capture its motion through video.
DNA nanotechnology takes advantage of the natural affinity for the DNA bases A, G, C and T to pair up with one another. By moving around the sequence of letters on synthetic strands of DNA, scientists can get the strands to bind together in ways that create different shapes and even build functioning machines.
The Salaita lab, a leader in biophysics and nanotechnology, developed the first rolling DNA-based motor in 2015. The device was 1,000 times faster than any other synthetic motor, fast-tracking the burgeoning field of molecular robotics. Its high speed allows a simple smart phone microscope to capture its motion through video.
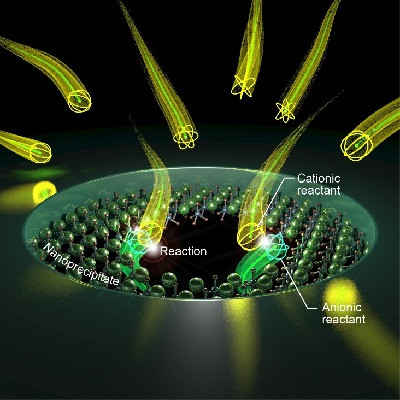
The motor’s “chassis” is a micron-sized glass sphere. Hundreds of DNA strands, or “legs” are allowed to bind to the sphere. These DNA legs are placed on a glass slide coated with the reactant RNA, the motor’s fuel. The DNA legs are drawn to the RNA, but as soon as they set foot on it they erase it through the activity of an enzyme that is bound to the DNA and destroys only RNA. As the legs bind and then release from the substrate, they keep guiding the sphere along.
When Piranej joined the Salaita lab in 2018, she began working on a project to take the rolling motors to the next level by building in computer programming logic.
“It’s a major goal in the biomedical field to take advantage of DNA for computation,” Piranej says. “I love the idea of using something that’s innate in all of us to engineer new forms of technology.”
DNA is like a biological computer chip, storing vast amounts of information. The basic units of operation for DNA computation are short strands of synthetic DNA. Researchers can change the “program” of DNA by tweaking the sequences of AGTC on the strands.
“Unlike a hard, silicon chip, DNA-based computers and motors can function in water and other liquid environments,” Salaita says. “And one of the big challenges in fabricating silicon computer chips is trying to pack more data into an ever-smaller footprint. DNA offers the potential to run many processing operations in parallel in a very small space. The density of operations you could run might even go to infinity.”
Synthetic DNA is also biocompatible and cheap to make. “You can replicate DNA using enzymes, copying and pasting it as many times as you want,” Salaita says. “It’s virtually free.”
Limitations remain, however, in the nascent field of DNA computation. A key hurdle is making the output of the computations easily readable. Current techniques heavily rely on tagging DNA with fluorescent molecules and then measuring the intensity of emitted light at different wavelengths. This process requires expensive, cumbersome equipment. It also limits the signals that can be read to those present in the electromagnetic spectrum.
Although trained as a chemist, Piranej began learning the basics of computer science and diving into bioengineering literature to try to overcome this hurdle. She came up with the idea of using a well-known reaction in bioengineering to perform the computation and pairing it with the motion of the rolling motors.
The reaction, known as toehold-mediated strand displacement, occurs on duplex DNA — two complementary strands. The strands are tightly hugging one another except for one loose, floppy end of a strand, known as the toe hold. The rolling motor can be programmed by coating it with duplex DNA that is complementary to a DNA target — a sequence of interest. When the molecular motor encounters the DNA target as it rolls along its RNA track, the DNA target binds to the toe hold of the duplex DNA, strips it apart, and anchors the motor into place. The computer read out becomes simply “motion” or “no motion.”
“When I first saw this concept work during an experiment, I made this really loud, excited sound,” Piranej recalls. “One of my colleagues came over and asked, ‘Are you okay?’ Nothing compares to seeing your idea come to life like that. That’s a great moment.”
These two basic logic gates of “motion” or “no motion” can be strung together to build more complicated operations, mimicking how regular computer programs build on the logic gates of “zero” or “one.”
Piranej took the project even further by finding a way to pack many different computer operations together and still easily read the output. She simply varied the size and materials of the microscopic spheres that form the chassis for the DNA-based rolling motors. For instance, the spheres can range from three to five microns in diameter and be made of either silica or polystyrene. Each alteration provides slightly different optical properties that can be distinguished through a cell phone microscope.
The Salaita lab is working to establish a collaboration with scientists at the Atlanta Center for Microsystems Engineered Point-of-Care Technologies, an NIH-funded center established by Emory and GeorgiaTech. They are exploring the potential for the use of the DNA-computing technology for home diagnostics of COVID-19 and other disease biomarkers.
“Developing devices for biomedical applications is especially rewarding because it’s a chance to make a big impact in people’s lives,” Piranej says. “The challenges of this project have made it more fun for me,” she adds.
Read the original article on eScienceCommons.