The researchers adapted the Broad Institute’s PRISM platform, which is designed to rapidly screen drugs on different cancer types, to cell-nanoparticle interactions to show if biological differences between cells explain why the effects of nanoparticles vary between indications. Using the resulting platform, the researchers barcoded 488 cancer cell lines from 22 different tissues of origin.
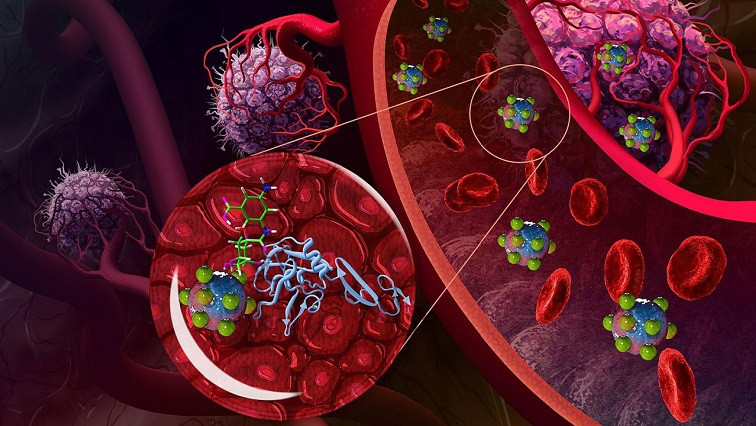
Working with the Broad Institute scientists behind PRISM, the researchers exposed hundreds of different cells to one of 35 fluorescently tagged nanoparticles made of liposomes or the polymers PLGA and polystyrene and coated with various protective and targeting molecules. The fluorescent tags enabled the researchers to sort the cells and assign each line a score based on its affinity for each nanoparticle.
“We are excited by our findings because it is really just the beginning—we can use this approach to map out what types of nanoparticles are best to target certain cell types, from cancer to immune cells and other kinds of healthy and diseased organ cells. We are learning how surface chemistry and other material properties play a role in targeting,” Paula Hammond, an MIT Institute Professor and co-author of the paper, said in a statement.
The findings, details of which were published in Science, included the identification of SLC46A3 as a protein deserving of further study. High levels of the protein correlated to very low uptake of lipid-based nanoparticles in the PRISM screen, a finding that was validated in a melanoma mouse model. If seen in humans, the correlation could help physicians identify patients who are likely to respond to treatment.
Read the original article on Fierce Pharma.