Previously, nanometer resolution had only been a theoretical possibility. Antibodies carrying a fluorescent dye to label cell structures could not reach their intended target.
In ExM, the sample to be examined is cross-linked into a swellable polymer. Then the interactions of the molecules in the sample are destroyed, and the sample is allowed to swell in water. This leads to an expansion: The molecules to be imaged drift apart spatially by a factor of four.
Previous attempts to combine the technique had been unsuccessful as fluorescent label dyes did not survive the polymerization process. Buffer solutions are also necessary for dSTORM, though the solutions cause the sample to shrink down to its original size.
“By stabilizing the gel and immune staining only after expansion, we could overcome these hurdles and successfully combine the two microscopy methods,” Sauer said. As a result, the distance error melts to just 5 nm when expanded 3.2× , which makes fluorescence imaging with molecular resolution possible.
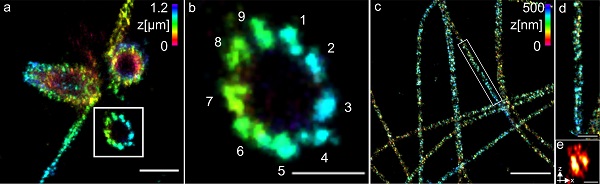
The researchers used centrioles and structures composed of the protein tubulin to demonstrate the method. They were able to visualize tubulin tubes as hollow cylinders with a diameter of 25 nm, and succeeded in imaging groups of three at a distance of 15 to 20 nm.
The team plans to apply the method to different structures, organelles, and multiprotein complexes of the cell. The study was carried out by researchers from the University of Würzburg, Monash University, and the University of Geneva.
The research was published in Nature Communications (www.doi.org/10.1038/s41467-020-17086-8).

